
Flow Cytometry Software
Frank Battye - Flow Cytometry Consulting
ACluE: For Flow Cytometry Data Cluster Analysis
New and
extensively revised for modern data clustering. Current New
Version ACluE3.0.3 released
28th December 2021.
[See also the new Weasel program for flow cytometry data analysis and display, upgraded to better support ACluE data files].
What ACluE does:
Haven't got a clue where your cell populations are? ACluE (Agglomerative Cluster Engine) is a cluster analysis program designed to search through flow cytometry data files seeking clusters of cells in mutual proximity within the multidimensional data space.
How ACluE does it:
The program relies on the ordered density approach of Goad [1], modified to use a mutual near neighbour value as the distance metric in order to better discriminate less dense clusters in proximity to denser clusters. See Gowda and Krishna [2]. Since the Goad algorithm requires a measure of the distance between each cell and every other, it is quite time consuming in execution. An important innovation in this revised version is therefore to use the Goad method to first discover "seed" clusters for a subset of the cellular data and thence to assign each further cell to the seed cluster from which it has sufficiently close Mahalanobis distance.
Having assigned each cell a cluster number, the program rewrites the flow cytometry data file with cluster number as an additional parameter. The cluster parameter can be used for colour coding (see below) or gating.
Note that ACluE knows nothing of flow cytometry; it knows nothing about fluorescent markers and treats cell data as points in a multi-dimensional space. The discovered clusters are therefore autonomous; not hierarchical.
Output:

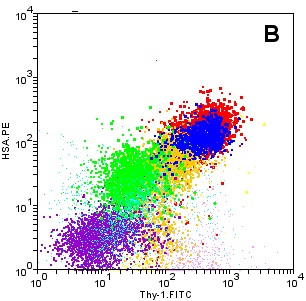
Early ACluE applications: A: Mature cell
types in lymph node have generally distinct populations so that
finding the clusters is not difficult. The display is a 7-axis
Biplot of the kind used in the Weasel program. B: Interpreting
data on murine thymic precursors (a cluster of which is coloured
green at display centre) among a collection of less mature cells
is not so straightforward. Clustering was determined on forward
scatter, side scatter and intensities of Thy-1, HSA, Class II
and autofluorescence. Although discoverable using ACluE,
the precursor cluster cannot be isolated on any 2D
dotplot. Data provided by Dr. Li Wu.
This full version of the new ACluE is available for testing by interested flow cytometrists. Comments and suggestions are solicited. Please download a copy. As has always been the case, ACluE is available free for permanent use to Weasel licence holders.
[1] C A Goad: Los Alamos Scientific Laboratory Biological & Medical Research Group Annual report LA-7120-MS 1978
[2] K C Gowda and G Krishna: Pattern Recognition 10 p105-112 1978
